A while back I got help on this issue I was having. This was the output that I was looking for:
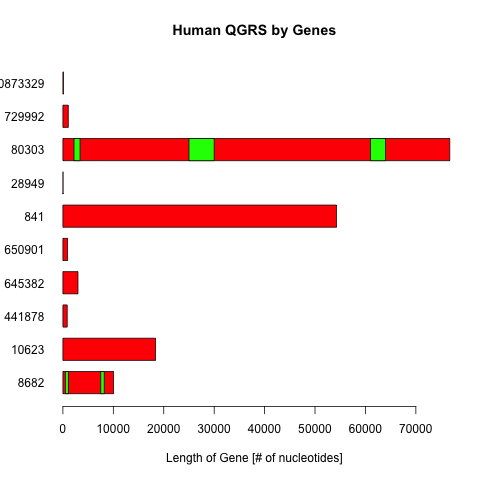
What it shows:
- Shows length of all chromosomes
- Shows position of a qgrs on a chromosome [multiple qgrs on a chromosome]
However, when I needed to tweak the data to work for what I needed, things just stopped working and I haven't been able to figure out what the problem is.
See my script in reply below, was having technical issues
.
.
.
.
.
.
.
Here is my data
For chr_plot:
248956422 1
242193529 2
198295559 3
190214555 4
181538259 5
170805979 6
159345973 7
145138636 8
138394717 9
133797422 10
135086622 11
133275309 12
114364328 13
107043718 14
101991189 15
90338345 16
83257441 17
80373285 18
58617616 19
64444167 20
46709983 21
50818468 22
156040895 23
57227415 24
For gq_plot:
389 407 1
524 547 1
739 755 1
834 861 1
904 927 1
942 969 1
1022 1051 1
1111 1129 1
1288 1303 1
1304 1327 1
1599 1628 1
1893 1903 1
1914 1924 1
1931 1952 1
1963 1979 1
2015 2033 1
2152 2179 1
2206 2225 1
2255 2277 1
2356 2383 1
2398 2415 1
2459 2472 1
2492 2506 1
2722 2740 1
3040 3062 1
3063 3082 1
3195 3214 1
3749 3777 1
3918 3937 1
4474 4501 1
4859 4873 1
4876 4889 1
4948 4964 1
5127 5155 1
5280 5309 1
5429 5439 1
6203 6232 1
6363 6391 1
6471 6488 1
6563 6589 1
6627 6652 1
7126 7155 1
7199 7227 1
7348 7374 1
7392 7421 1
7432 7442 1
7495 7507 1
7559 7568 1
7774 7803 1
7875 7887 1
8021 8044 1
8297 8307 1
8792 8813 1
8825 8854 1
8866 8886 1
8971 8986 1
9011 9028 1
9076 9104 1
9145 9168 1
9358 9377 1
9647 9659 1
10098 10118 1
10158 10173 1
10351 10361 1
10474 10494 1
10498 10519 1
10800 10823 1
11720 11729 1
12275 12304 1
12531 12547 1
12642 12666 1
12818 12845 1
12878 12900 1
13104 13124 1
13175 13199 1
13640 13667 1
14244 14259 1
14389 14417 1
14908 14924 1
14992 15010 1
15126 15148 1
15224 15253 1
15269 15292 1
15385 15413 1
15782 15804 1
15987 16016 1
16545 16559 1
16587 16615 1
16724 16739 1
16756 16767 1
16885 16906 1
17157 17186 1
17864 17874 1
17876 17886 1
17944 17973 1
18063 18078 1
18101 18126 1
18139 18167 1
18237 18264 1
18344 18362 1
18542 18564 1
18607 18630 1
18772 18801 1
18812 18839 1
19028 19045 1
19073 19102 1
19105 19121 1
19696 19724 1
19849 19875 1
21247 21276 1
21719 21746 1
21942 21966 1
22053 22070 1
22096 22111 1
22197 22210 1
22232 22247 1
22403 22419 1
22635 22664 1
22754 22783 1
22827 22847 1
22977 22998 1
23015 23043 1
23340 23358 1
23431 23443 1
24586 24614 1
25070 25099 1
25131 25153 1
25180 25195 1
25279 25293 1
25316 25331 1
25349 25359 1
25624 25638 1
25863 25879 1
25905 25920 1
26074 26098 1
26331 26355 1
26393 26413 1
26439 26453 1
26592 26620 1
27028 27053 1
27395 27409 1
29061 29075 1
29106 29121 1
29211 29234 1
29247 29262 1
29582 29600 1
29729 29743 1
29895 29910 1
30028 30043 1
30230 30257 1
30461 30485 1
30867 30893 1
31015 31039 1
31538 31563 1
31766 31793 1
31917 31935 1
32298 32325 1
32326 32345 1
33090 33105 1
33134 33148 1
33174 33189 1
33316 33331 1
33350 33360 1
33728 33745 1
33772 33786 1
33910 33924 1
34369 34395 1
35018 35032 1
35059 35074 1
35160 35173 1
35196 35211 1
35592 35615 1
37128 37154 1
37242 37267 1
37634 37661 1
38841 38869 1
39168 39182 1
39306 39320 1
39638 39648 1
39726 39745 1
39805 39829 1
39917 39937 1
39959 39988 1
40000 40025 1
40139 40154 1
40318 40343 1
40484 40510 1
40611 40638 1
40681 40709 1
40733 40760 1
40834 40861 1
40937 40965 1
41111 41140 1
41159 41178 1
41218 41231 1
41273 41296 1
41325 41347 1
41383 41408 1
42097 42123 1
42231 42245 1
42280 42296 1
42422 42448 1
42665 42686 1
43560 43579 1
43960 43988 1
1137 1176 1
1842 1885 1
2045 2083 1
2093 2124 1
2957 3001 1
3279 3317 1
6655 6680 1
7303 7347 1
7719 7759 1
10298 10329 1
11316 11346 1
11472 11500 1
12068 12109 1
14852 14888 1
14930 14964 1
17545 17560 1
38509 38534 1
40896 40924 1
42033 42077 1
Please note that the gq_plot data is everything I will be using for my project, but only for chromosome 1. I still have all the other 23 chromosomes and data for them. And I need to be able to use this script to show all of that in one!
So I will be putting all the other "qgrs" data for the other chromosomes in this same file. So hopefully it will just work with the script once I can get the script working.
Now obviously there is so much data, and many of the qgrs lengths are very small. So we are all wondering, how are we going to see any of this??? If nothing really comes up because it's that bad, then I thought I could either have a single graph generated per chromosome, instead of one with all 24 chromosomes, OR even better, I could have some kind of user defined zoom function.
It would be simple, basically the user gets to set the length they view.
But I don't know how to do that... so any help would be great! I've tried for months, so at this point I am not really looking for advice, just a completed project. It would mean so much to me! THANKS!!